gold序列实验报告Word文档格式.docx
《gold序列实验报告Word文档格式.docx》由会员分享,可在线阅读,更多相关《gold序列实验报告Word文档格式.docx(15页珍藏版)》请在冰豆网上搜索。
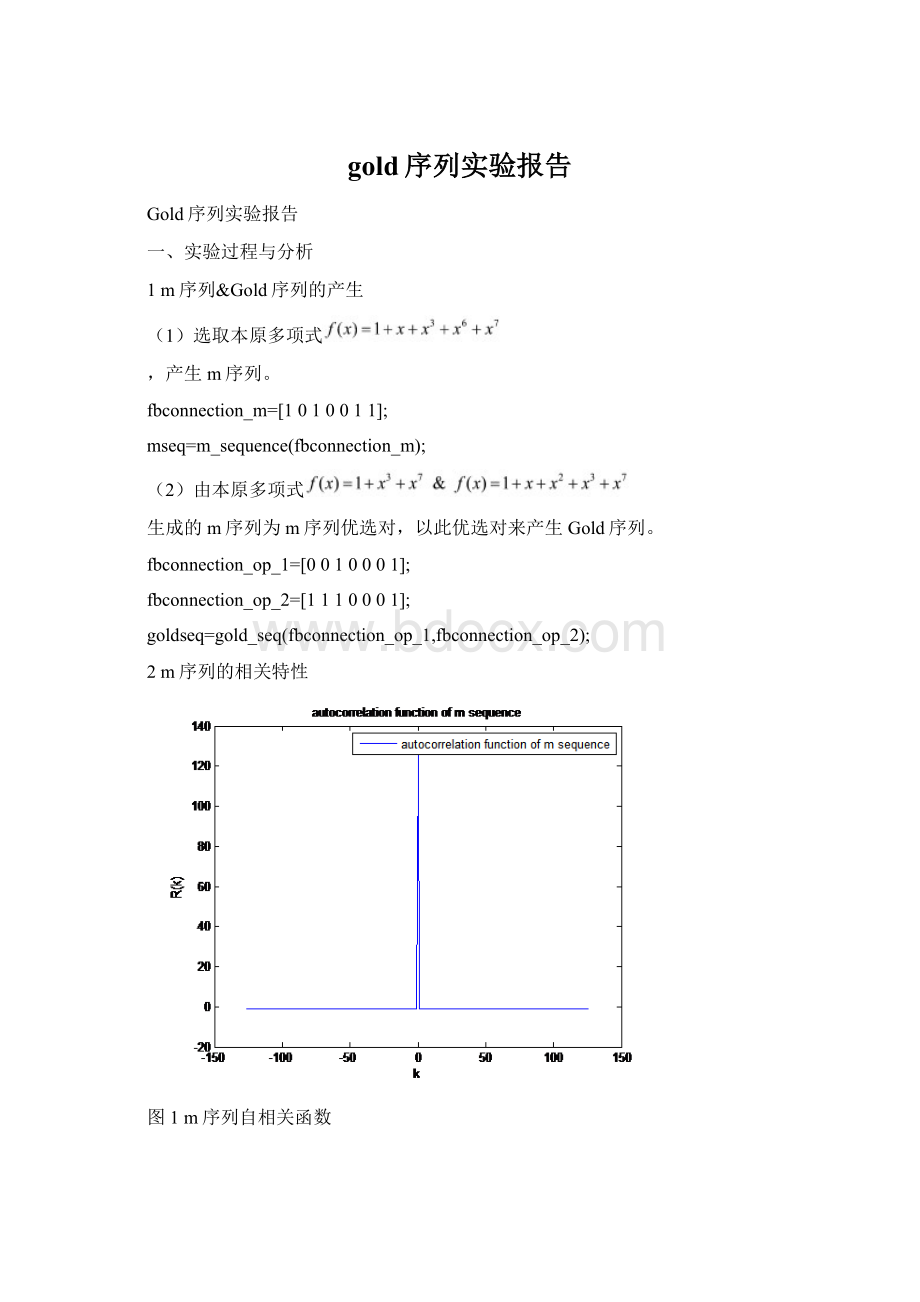
Gold序列相关特性比较
图6m序列&
Gold序列自相关函数比较
图7m序列&
Gold序列互相关函数比较
gold序列在时延不为0是三值的,这一点不如m序列,在时延为0处具有与m序列相同的峰值特性。
显然,m序列的自相关特性比Gold序列的自相关特性要好。
Gold序列具有更小的互相关峰值,显然Gold序列的互相关特性比m序列的互相关特性性能要好。
5m序列优选对
m序列优选对也是gold序列族中的gold序列,所以其互相关函数与gold序列一致。
图8m序列优选对的互相关函数
6任选6个Gold序列,观察其自相关函数
图9任选6个Gold序列的自相关函数
7任选6对Gold序列,观察其互相关函数
图10任选6对Gold序列的互相关函数
任意两个Gold序列互相关函数幅度小于或等于17,符合Gold理论。
源码
(1)PN_Correlation.m
%=======================================
%PN_Correlation.m
%CorrelationanalysisofPNsequence
%
%Author:
jzh(04011646seu)
%clean
closeall;
clearall;
clc;
%optimumpairs
%anymsequencefortest
%goldsequence
mseq_op_1=m_sequence(fbconnection_op_1);
mseq_op_2=m_sequence(fbconnection_op_2);
%msequence
%mapping
mseq_op_1((mseq_op_1==0))=-1;
mseq_op_2((mseq_op_2==0))=-1;
goldseq((goldseq==0))=-1;
mseq((mseq==0))=-1;
m_auto=ccorr(mseq);
%anytwomsequencesexceptoptimumpairs
m_cross=ccorr(mseq,mseq_op_1);
gold_auto=ccorr(goldseq(1,:
));
gold_cross=ccorr(goldseq(1,:
),goldseq(2,:
m_op_cross=ccorr(mseq_op_1,mseq_op_2);
m_auto_max=max(m_auto)
m_auto_min=min(m_auto)
gold_auto_max=max(gold_auto)
m_cross_amplitude_max=max(abs(m_cross))
gold_cross_amplitude_max=max(abs(gold_cross))
N=length(mseq);
%sequencelength
abscissa=-N+1:
N-1;
%abscissaaxis
figure
(1);
plot(abscissa,m_auto);
xlabel('
k'
);
ylabel('
R(k)'
legend('
autocorrelationfunctionofmsequence'
title('
figure
(2);
plot(abscissa,m_cross);
cross-correlationfunctionofmsequence'
figure(3);
plot(abscissa,gold_auto);
autocorrelationfunctionofgoldsequence'
figure(4);
plot(abscissa,gold_cross);
cross-correlationfunctionofgoldsequence'
figure(5);
subplot(1,2,1);
subplot(1,2,2);
figure(6);
figure(7);
plot(abscissa,m_op_cross);
cross-correlationfunctionofmsequenceoptimumpairs'
%testautocorrelationofanygoldsequence
fortest_time=1:
6
ii=ceil(rand()*(N+2));
figure();
temp_gold_auto=ccorr(goldseq(ii,:
plot(abscissa,temp_gold_auto);
xlabel('
legend(strcat('
seqnumber='
num2str(ii)));
title('
autocorrelationfunctionofarandomgoldsequence'
end
%testcross-correlationofany2goldsequence
jj=ceil(rand()*(N+2));
whilejj==ii
end
temp_gold_cross=ccorr(goldseq(ii,:
),goldseq(jj,:
plot(abscissa,temp_gold_cross);
seq1number='
num2str(ii),'
seq2number='
num2str(jj)));
title(strcat('
cross-correlationfunctionof2randomgoldsequence:
'
...
'
crosscorrelationamplitudeMAX='
num2str(max(abs(temp_gold_cross)))));
(2)m_sequence.m
%==============================
%m_sequence.m
%generatingmsequence
functionmseq=m_sequence(fbconnection)
%==============================m_sequence==============================
%Input:
%fbconnection:
feedbackconnectionrespondingtoprimitivepolynomial
%Output:
%msequ:
generationofmsequence,1-by-N
%========================================================================
n=length(fbconnection);
N=2^n-1;
%lengthofmsequence
register=[zeros(1,n-1)1];
%initialstateofshiftregister
%preallocating
newregister=zeros(1,n);
mseq=zeros(1,N);
mseq
(1)=register(n);
forii=2:
N
%feedback
newregister
(1)=mod(sum(fbconnection.*register),2);
%shift
forjj=2:
n
newregister(jj)=register(jj-1);
register=newregister;
%update
mseq(ii)=register(n);
%outputmsequence
(3)gold_seq.m
%==================================
%gold_seq.m
%generatinggoldsequencefamily
functiongoldseq=gold_seq(fbconnection1,fbconnection2)
%===============================gold_seq===============================
%fbconnection1:
feedbackconnection1
%fbconnection2:
feedbackconnection2
%Note:
fbconnection1&
fbconnection2areoptimumpairs
%goldseq:
generationofgoldsequencefamily,(N+2)-by-N
mseq1=m_sequence(fbconnection1);
mseq2=m_sequence(fbconnection2);
N=2^length(fbconnection1)-1;
goldseq=zeros(N+2,N);
%preallocating
%differentshiftamountoftwomsequence
forshift_amount=0:
N-1
shift_mseq2=[mseq2(shift_amount+1:
N)mseq2(1:
shift_amount)];
goldseq(shift_amount+1,:
)=mod(mseq1+shift_mseq2,2);
goldseq(N+1,:
)=mseq1;
goldseq(N+2,:
)=mseq2;
(4)ccorr.m
%ccorr.m
%calculatecross-correlationfunction
functionr=ccorr(seq1,seq2)
%================================ccorr=================================
%seq1:
sequence1
%seq2:
sequence2
(a)lengthoftwosequencemustbethesame
%(b)twoinputsequence:
calculatecross-correlationfunction
%(c)oneinputsequence:
calculateautocorrelationfunction
%r:
correlationfunction,1-by-2*N-1
%oneinputsequence:
calculateautocorrelation
ifnargin==1
seq2=seq1;
N=length(seq1);
r=zeros(1,2*N-1);
fork=-N+1:
-1
seq2_shift=[seq2(k+N+1:
N)seq2(1:
k+N)];
r(N+k)=seq1*seq2_shift'
;
fork=0:
seq2_shift=[seq2(k+1:
k)];